By Caio Figueiredo, Veterinary Medicine Extension
Bovine respiratory disease (BRD) complex remains one of the most common and costly livestock diseases worldwide. Despite the existence of almost 11,000 publications related to BRD (the first from the early 1900’s) in PubMed (accessed on 07/10/25), the mechanisms underlying BRD occurrence and the best management practices for affected animals are not completely known. In the U.S., BRD is estimated to affect approximately 22% of dairy calves, and is associated with 30% of all calf mortalities (Dubrovsky et al., 2019). Aside from the animal welfare issues and economic losses associated with BRD, the increased use of antimicrobials represents another challenge. Respiratory diseases are the 2nd highest reason for antimicrobial use in pre-weaned heifers, and approximately 95% of affected animals receive antimicrobials (USDA, 2018). Considering the importance of BRD for dairy sustainability, and possibly public health, the main objective of this article is to highlight recently published manuscripts related to the use of antimicrobials, and genomic selection for the management of BRD in dairy cattle.
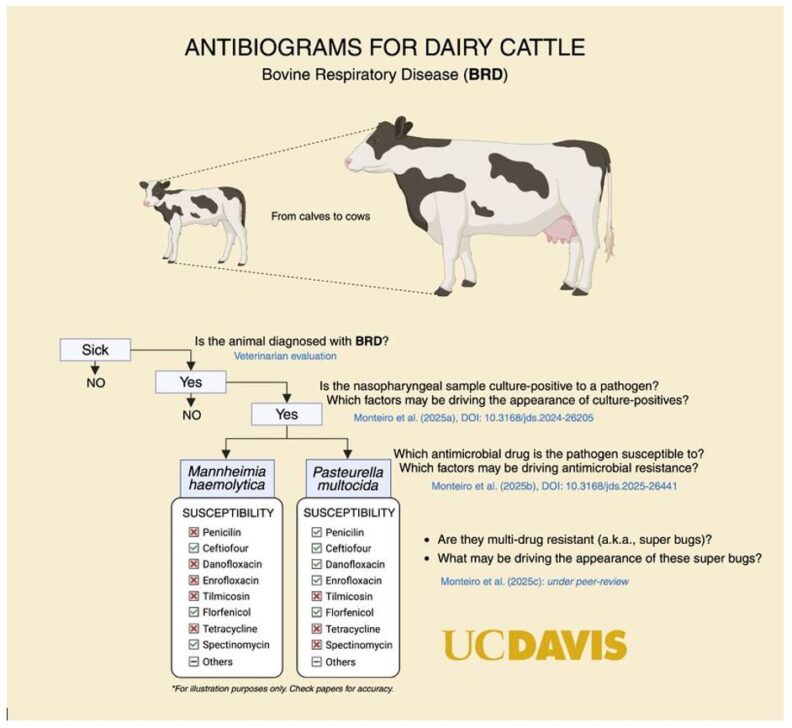
Dr. Richard Pereira and collaborators from UC Davis published a series of papers that focused on several aspects of antibiogram use for BRD in dairy cattle. One of the main challenges for BRD management is the limited information available regarding factors associated with herd-level bacterial pathogen recovery for BRD clinical cases. That information is important to help identify strategies to support the successful collection of a minimum number of each organism over time to generate cumulative antibiogram susceptibility testing reports. For their study (Monteiro et al., 2025a), samples were collected from a total of 301 BRD clinical cases: 146 samples with a culture-positive for P. multocida, 63 samples with a culture-positive for M. haemolytica, and 3 samples with a culture-positive for Histophilus somni. For factors associated with the culture-positive P. multocida BRD clinical cases, an interaction between age and season was identified, where cows had overall lower odds of being culture-positive independently of the season when compared with calves in the spring and summer and heifers in the fall and winter. For factors associated with the culture-positive M. haemolytica BRD clinical cases, an interaction was also observed between age and season, but the farm further played a role in the odds of being culture-positive, with one farm having considerably greater odds than the remaining ones.
A subsequent study aimed to evaluate a cumulative antimicrobial susceptibility testing framework for BRD in dairy cattle and factors of relevance when analyzing and reporting antibiogram data for on-farm treatment decisions. In that study (Monteiro et al., 2025b), calves, heifers, and cows from 3 dairy farms in California were sampled using deep nasopharyngeal swabs for culture and antimicrobial susceptibility testing for P. multocida and M. haemolytica. Antimicrobial susceptibility testing was performed using the minimum inhibitory concentration approach for 12 antimicrobial drugs of relevance to dairy cattle. The variability in the odds of unsusceptible (resistant) isolates was evaluated for effects of age, farm, and season. Overall, P. multocida isolates were primarily resistant to tetracycline (74%) and spectinomycin (46%), whereas M. haemolytica isolates were frequently resistant to penicillin (43%) and tetracycline (34%). Calves and heifers showed a higher number of resistant isolates than cows. For P. multocida specifically, isolates resistant to tetracycline and spectinomycin were affected by age group, with calves showing greater odds of resistance. A farm effect was significantly associated with P. multocida resistance to all antimicrobials except penicillin, suggesting individual farm management practices may heavily affect profiles of bacterial resistance. Similarly, M. haemolytica resistance was consistently associated with farm for macrolides and tetracyclines, and heifers showed the greatest odds of resistance when compared to cows. Season did not affect the resistance of either pathogen, suggesting that California’s seasonal patterns or other factors, such as farm practices and animal management, play a more prominent role in susceptibility profiles.
Because of concerns related to the creation of multidrug-resistant (MDR) microorganisms, a third study aimed to characterize the MDR profile of P. multocida and M. haemolytica isolates from the deep nasopharyngeal swabs of calves, heifers, and cows diagnosed with BRD (Monteiro et al., 2025c). Overall, there was a low prevalence of MDR in both P. multocida and M. haemolytica in the study, but farm management and the age of the animals were associated with MDR prevalence. Altogether, the recently published studies shed light on possible ways to manage BRD in dairy cattle (including antibiogram use; Figure 1), factors associated with microorganism collection and culture, and possible factors tied to MDR microorganisms.
This brings us to the use of genomic selection which has intensified over the years and prompted the development of different indexes that include genomic regions related to productive performance and animal health. Some of our own researchers at WSU have made valuable contributions that highlight the genetic differences associated with BRD, and that could lead to improved animal selection models. For example, a recent publication from Dr. Holly Neibergs’ research group and collaborators provided insight and validation of genomic regions that enhance selection for BRD resistance and for healthier cattle. Herrick et al. (2025) reported that over 62 loci and 123 positional candidate genes were associated with BRD in pre-weaned calves. In addition, over 181 loci and 185 positional candidate genes were associated with BRD in post-weaned calves. Moreover, our Field Disease Investigation Unit has reported preliminary data related to differences in immunoregulation in transition cows associated with previous BRD occurrence (Richmond et al., Abstract #1251). Overall, reported immunoregulatory differences in lactating dairy cows suggest that preweaning disease may have underlying and persistent impacts on immune factors influencing the periparturient period. Further exploration of these factors may provide a complementary insight into associated disease phenotypes and transition cow management. Additionally, this can further inform genetic selection strategies aimed at reducing production losses while increasing disease resiliency. Although there is much to be discovered relative to BRD, recent research shares promising data that could further help improve judicious use of antimicrobials and reduce the occurrence of BRD by selecting less susceptible animals.